Puzzle-solving: More Than Just a Game
We have another guest post from our Communications Research Executive Naomi! Check out her other RFS blog post here if you haven’t read it.
Puzzle-solving: More Than Just a Game
Codes, ciphers, mysteries and puzzles – it all seems a bit Tom Hanks running around in the invariably sunny Italian setting of the latest Dan Brown adaptation.
Well, Cambridge has beautiful historic buildings, and an intriguing history of spies and secrets. We can’t promise sunshine, but we can guarantee that Race For Science will be a lot of fun.
But solving puzzles is an important part of life, and something we’ve always, as humans, tried to do. Just look at the Imago Mundi, the world’s earliest known map, dated to 500 BCE and discovered in the Iraqi city of Sippar, just north of the ancient city of Babylon. The map puts Babylon at the centre, and while many real mountains, cities and rivers are engraved on the clay tablet, so are imaginary, mystical features. This isn’t uncommon in early maps, where monsters and mythical creatures lurked around the edges of chartered territory. People put down what they know about the world and falling back on what they think they know to plug the gaps.

To the best of my knowledge, there are no dragons or sea monsters at Race For Science (I’ll double check with Jess), but using a similar thought process is helpful when you’ve got a clue to solve quickly – think first about what you know, and use that as a basis for your wildest guesses!
I would say that puzzle-solving is also at the core of cancer research, like the research that Race For Science is funding.
For example, when I did my PhD, I spent a lot of time investigating the PI3K pathway. Believe it or not, this picture below is a highly simplified version of it. This is a kind of emergency pathway some breast cancer cells could switch on when a common drug called Herceptin was used. We were trying to block this pathway, so that Herceptin would keep killing breast cancer cells and patients wouldn’t have their cancer come back. But if you block a single protein, it’s like putting a roadblock in front of a car being driven by a particularly belligerent motorist. It will try to plough through it. If it can’t do that, it will try to go around it, usually by finding other proteins to switch on when a drug turns a protein off.
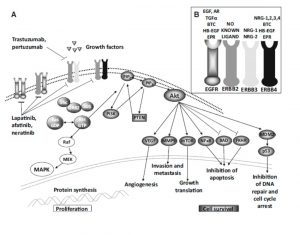
Simplified(!) PI3K Pathway. Reference: Elster et al, Breast Cancer Research and Treatment, 2014
Luckily, scientists have come up with many clever concepts to shut down this kind of drug resistance (the study I was part of led to a clinical trial). Science is highly detailed. But looking at the big picture, it’s not unlike tracking your way through a maze.
A lot of the work which happens in biology labs is abstract. In a common technique called a western blot, for example, you spend a few days and then get a band to appear on a piece of film. The band tells you a specific protein is present in whatever you have been working with, whether that was a piece of a real tumour a patient had donated, a piece of non-cancerous tissue, or cells artificially grown in the laboratory. You then have to think about what that protein might do, how there’s more or less of it in cancer than normal cells, and how that might change the behaviour of cancer. It’s a bit like those ancient maps – or filling in a crossword. You do research to find answers to unanswered questions, and when you get a clue or a new piece of information, you relate that to what’s already known to make sense of it.
Our PCRC scientists have important puzzles to solve. Christine is trying to train the immune system to fight cancer, while Magali is decoding how prostate cancer spreads, with a view to stopping it spreading beyond the prostate. Aamir is working out whether drugs used in other diseases can be “repurposed” as a new and low-cost treatment for cancer, and Matt is working on new prostate cancer lab models, kind of like decode keys which should speed up the process of getting new treatments to men with prostate cancer.
An immersive game and a real-life laboratory might seem like worlds apart. I’m going to be a bit cheeky and say that Race For Science strikes me as being a p-value of less than 0.05 more fun than most of the routine things I did in the lab (sorry, that’s a joke for the scientists – and admittedly not even a particularly good one). But at the end of the day, both are contributing to a better future for men with prostate cancer and the families of those men, and both are doing it by solving puzzles.

